Topic 4: Introduction to teal.modules.clinical and anatomy of a module (15 minutes)
What is teal.modules.clinical?
The teal.modules.clinical (TMC) package offers specialized modules designed specifically for clinical trial data analysis. These modules are built on top of teal.modules.general and provide clinical-specific functionality and workflows.
Launching a simple teal.modules.clinical application
Here’s a simple example using a teal.modules.clinical module:
library(teal)
library(teal.modules.clinical)
library(teal.data)
library(pharmaverseadam)
# Step 1: Load real ADaM data from pharmaverseadam
data("adsl", package = "pharmaverseadam")
adsl$ARM <- as.factor(adsl$ARM)
adsl$ARMCD <- as.factor(adsl$ARM)
# Step 2: Create teal_data object with the "real" dataset
clinical_data <- cdisc_data(
ADSL = adsl,
code = "
adsl$ARM <- as.factor(adsl$ARM)
adsl$ARMCD <- as.factor(adsl$ARM)
ADSL <- adsl
"
)
# Step 3: Create the teal application with a TMC module
app <- teal::init(
data = clinical_data,
modules = teal::modules(
teal.modules.clinical::tm_t_summary(
label = "Demographic Table",
dataname = "ADSL",
arm_var = choices_selected(c("ARM", "ARMCD"), "ARM"),
add_total = TRUE,
summarize_vars = choices_selected(
c("SEX", "RACE", "BMRKR2", "EOSDY", "DCSREAS", "AGE"),
c("SEX", "RACE")
),
useNA = "ifany"
)
)
)
# Step 4: Launch the application
if (interactive()) {
shiny::shinyApp(app$ui, app$server)
}Anatomy of a teal.modules.clinical module
When you launch a teal.modules.clinical application, you’ll observe:
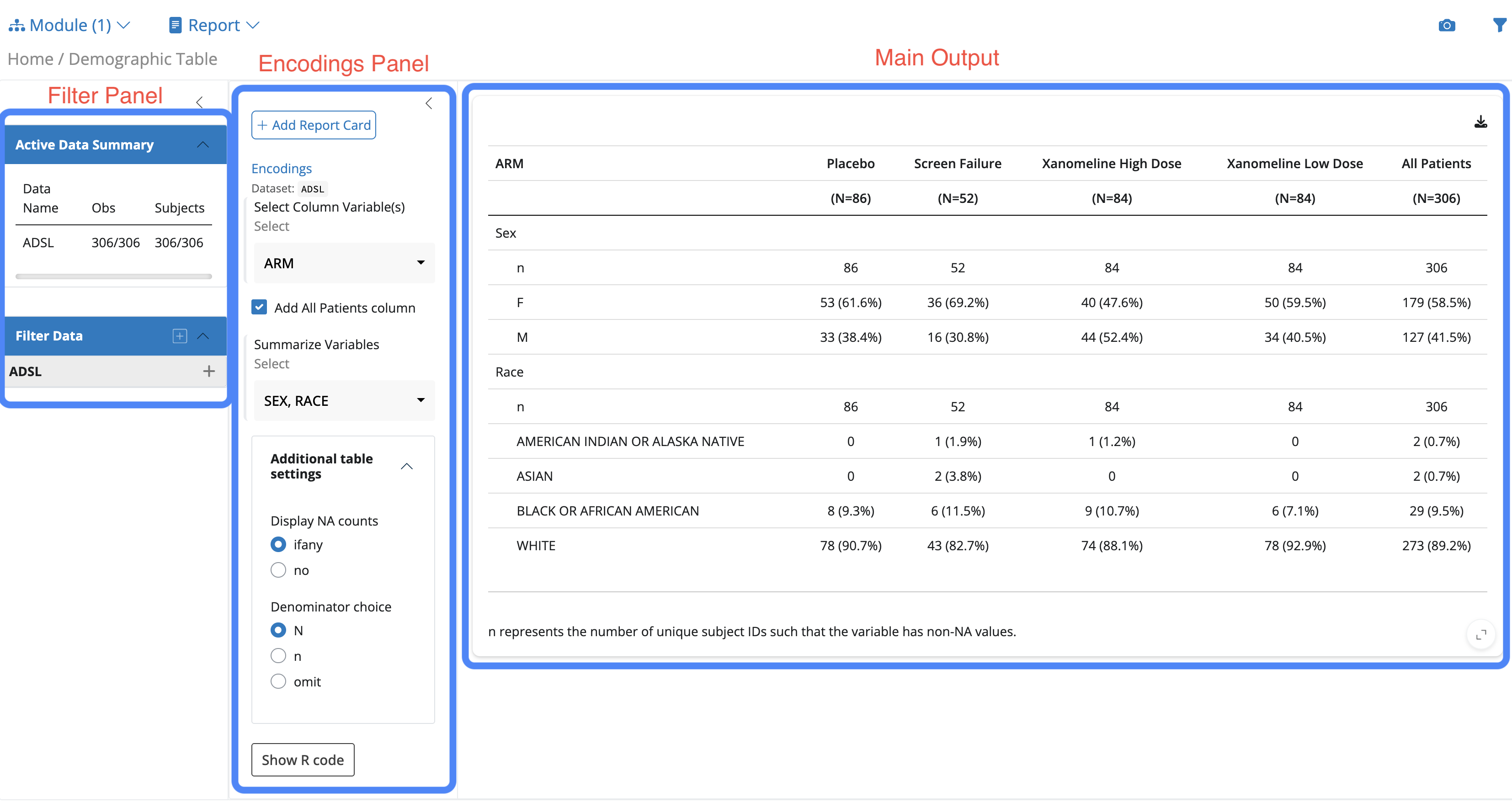
teal module- Filter Panel (Left): Shows filtering options for all dataset variables
- Categorical variables: Display as checkboxes (e.g., SEX, ARM, RACE)
- Numeric variables: Display as range sliders (e.g., AGE, BMIBL)
- Date variables: Display as date pickers (e.g., RFSTDTC, RFENDTC)
- All filters are interconnected with module outputs
- Encoding Panel: Provides controls for configuring the analysis
- Variable selection dropdowns
- Analysis parameter settings
- Customization options specific to the module type
- Show R Code Button: generates reproducible R code for all operations:
- Click to view the exact R code used
- Copy code for reproducibility
- Ensures transparency in analysis
- Main Output (Center): Displays the module’s primary analytical output
- Module-specific visualizations or tables
- Interactive elements for exploration
🛠️ Exercise
- create a
cdisc_dataobject usingADSLandADAEfrompharmaverseadam - use
join_keys(created_data)to ensure that your data has correctjoin_keys
🛠️ Exercise
- using documentation of
teal.modules.clinical- https://insightsengineering.github.io/teal.modules.clinical/latest-tag/reference/tm_t_events.html create atealapplication including thetm_t_eventsmodule - change the default sort criterium to alphanumeric
- add a title to the module’s output using
pre_output
TipAnswer
library(teal.data)
library(teal)
library(pharmaverseadam)
library(teal.modules.clinical)
ADSL <- pharmaverseadam::adsl
ADSL$ARM <- as.factor(ADSL$ARM)
ADSL$ARMCD <- as.factor(ADSL$ARMCD)
my_data <- cdisc_data(ADSL = ADSL, ADAE = pharmaverseadam::adae, code = "
ADAE <- pharmaverseadam::adae
ADSL <- pharmaverseadam::adsl
ADSL$ARM <- as.factor(ADSL$ARM)
ADSL$ARMCD <- as.factor(ADSL$ARMCD)
")
app <- init(
data = my_data,
modules = modules(
tm_t_events(
label = "Adverse Event Table",
dataname = "ADAE",
arm_var = choices_selected(c("ARM", "ARMCD"), "ARM"),
llt = choices_selected(
choices = variable_choices("ADAE", c("AETERM", "AEDECOD")),
selected = c("AEDECOD")
),
hlt = choices_selected(
choices = variable_choices("ADAE", c("AEBODSYS", "AESOC")),
selected = "AEBODSYS"
),
add_total = TRUE,
event_type = "adverse event",
sort_criteria = "alpha",
pre_output = shiny::div("Who won the most at the poker last night?")
)
)
)
shinyApp(app$ui, app$server)🛠️ Exercise (for all the early finishers)
- using documentation - https://insightsengineering.github.io/teal.modules.clinical/latest-tag/reference/tm_t_events_summary.html create a
tealapplication including thetm_t_events_summarymodule.
TipAnswer
library(dplyr)
library(teal.modules.clinical)
data <- teal_data()
data <- within(data, {
ADSL <- pharmaverseadam::adsl %>%
mutate(
DTHFL = case_when(
!is.na(DTHDT) ~ "Y",
TRUE ~ ""
) %>% with_label("Subject Death Flag")
)
ADSL$DCSREAS <- NA_character_
ADSL$ARM <- as.factor(ADSL$ARM)
ADSL$ARMCD <- as.factor(ADSL$ARMCD)
ADAE <- pharmaverseadam::adae
ADAE$ARM <- as.factor(ADAE$ARM)
ADAE$ARMCD <- as.factor(ADAE$ARMCD)
ADAE$DCSREAS <- NA_character_
levels(ADAE$ARM) <- levels(ADSL$ARM)
})
print(data)
print(names(data[["ADAE"]]))
join_keys(data) <- default_cdisc_join_keys[names(data)]
app <- init(
data = data,
modules = modules(
tm_t_events_summary(
label = "Adverse Events Summary",
dataname = "ADAE",
arm_var = choices_selected(
choices = variable_choices("ADSL", c("ARM", "ARMCD")),
selected = "ARM"
),
add_total = TRUE
)
)
)
shinyApp(app$ui, app$server)🌐 References
tealdocumentationteal.modules.generaldocumentationteal.modules.clinicaldocumentationteal.datadocumentation